![[*]](http://sepwww.stanford.edu/latex2html/cross_ref_motif.gif) )
that we invented earlier.
These can be thought of as the solution to a spectral
factorization problem:
)
that we invented earlier.
These can be thought of as the solution to a spectral
factorization problem:
When an image is spoiled by excessive low frequencies, we can cut them out by taking a gradient or a helix derivative.
Neither the gradient nor the helix derivative
has an adjustable cutoff frequency.
A slightly more complicated form allows us to
build a lowcut filter with an adjustable cut-off frequency.
Recall the one-dimensional causal lowcut filters (![[*]](http://sepwww.stanford.edu/latex2html/cross_ref_motif.gif) )
that we invented earlier.
These can be thought of as the solution to a spectral
factorization problem:
)
that we invented earlier.
These can be thought of as the solution to a spectral
factorization problem:
| (18) |
After some experimentation with this filter,
I learned that I already had a powerful adjustable
parameter in the number of coefficients
that are used from the spectral factorization of ![]() .I learned that taking not enough lags in the filter
caused it not to drop to zero at zero frequency.
A simple test of whether it drops to zero at zero frequency
is to add all the coefficients
in the definition (11)
and see how close they come to zero.
Doing this for the 8 coefficients given,
I find the sum is .18 compared to the lead coefficient 1.79,
so it seems the ``zero'' at zero frequency actually only drops 90%
of the way to zero.
Thus, although I coded a filter response to be the factorization of
.I learned that taking not enough lags in the filter
caused it not to drop to zero at zero frequency.
A simple test of whether it drops to zero at zero frequency
is to add all the coefficients
in the definition (11)
and see how close they come to zero.
Doing this for the 8 coefficients given,
I find the sum is .18 compared to the lead coefficient 1.79,
so it seems the ``zero'' at zero frequency actually only drops 90%
of the way to zero.
Thus, although I coded a filter response to be the factorization of
![]() ,I then abandoned it.
Instead, I choose the depth of the hole at zero frequency
by adjusting the number of filter coefficients
in the representation of the helix derivative.
,I then abandoned it.
Instead, I choose the depth of the hole at zero frequency
by adjusting the number of filter coefficients
in the representation of the helix derivative.
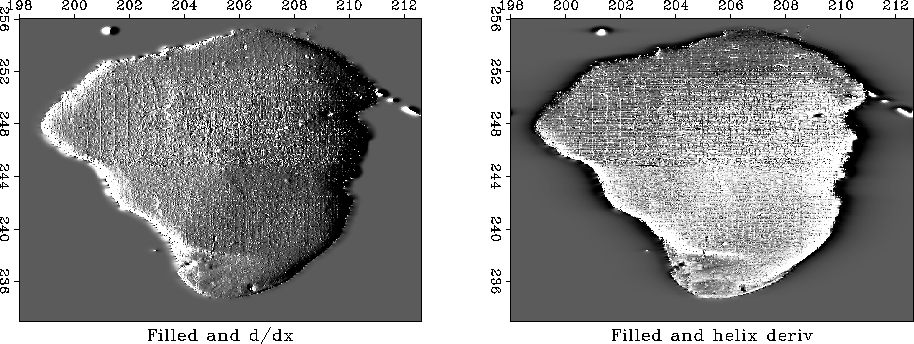 |
Figure 9 shows the application of the helix derivative to the Sea of Galilee. In experimenting with preparing this figure I found it best to use a filter with a whopping 128 points. With smaller numbers of points, the middle of the lake tended to be a dome (or bowl) just like the lake itself. The reason is that the helix derivative is not fully eliminating the zero frequency. If computation time were a burden, we might simply prefer to begin from a smaller helix derivative representation but then artificially reduce the size of the zero lag input (or boost the other values).
An operator for applying the helix filter is
helderiv ![[*]](http://sepwww.stanford.edu/latex2html/cross_ref_motif.gif) .
.
#$
#$=head1 NAME
#$
#$helderiv - helix derivative filter
#$
#$=head1 SYNOPSIS
#$
#$Initializer: C<call helderiv_init(n1,na)>
#$
#$Operator : C<ierr=helderiv_lop(adj,add,pp,qq)>
#$
#$Close : C<call helderiv_close()>
#$
#$=head1 PARAMETERS
#$
#$=over 4
#$
#$=item n1 - int
#$
#$ number of samples in n1
#$
#$=item na - int
#$
#$ number of coefs for filter
#$
#$=item adj - logcal
#$
#$ wheter or not run the adjoint
#$
#$=item add - logical
#$
#$ whether or not to add outout to data
#$
#$=item pp - C<real(:)>
#$
#$ model
#$
#$=item qq - C<real(:)>
#$
#$ data
#$
#$=back
#$
#$=head1 DESCRIPTION
#$
#$ Create and apply a helix derivative filter
#$
#$
#$=head1 SEE ALSO
#$
#$L<lapfac>, L<helicon>
#$
#$=head1 LIBRARY
#$
#$B<geef90>
#$
#$=cut
module helderiv {
use lapfac
use helicon
type( filter), private :: aa
#% _init( n1, na, eps)
integer, intent (in) :: n1, na
real, intent (in) :: eps
aa = lapfac2( eps, n1, na)
call helicon_init( aa)
#% _lop (pp, qq)
integer stat1
stat1 = helicon_lop( adj, .false., pp, qq)
#% _close
call deallocatehelix( aa)
}
Figure 10 shows the application of the helix derivative to a medical X-ray. Experimentation with this figure showed that the helix filter need not be long (16 points was enough) but, since the inputs were all positive numbers, the output was best displayed with its mean removed because the short 16 point filter did not remove the mean.
|
mam
Figure 10 Mammogram (medical X-ray). The white circles are metal foil used for navigation. The cancer is the ``spoked wheel.'' | 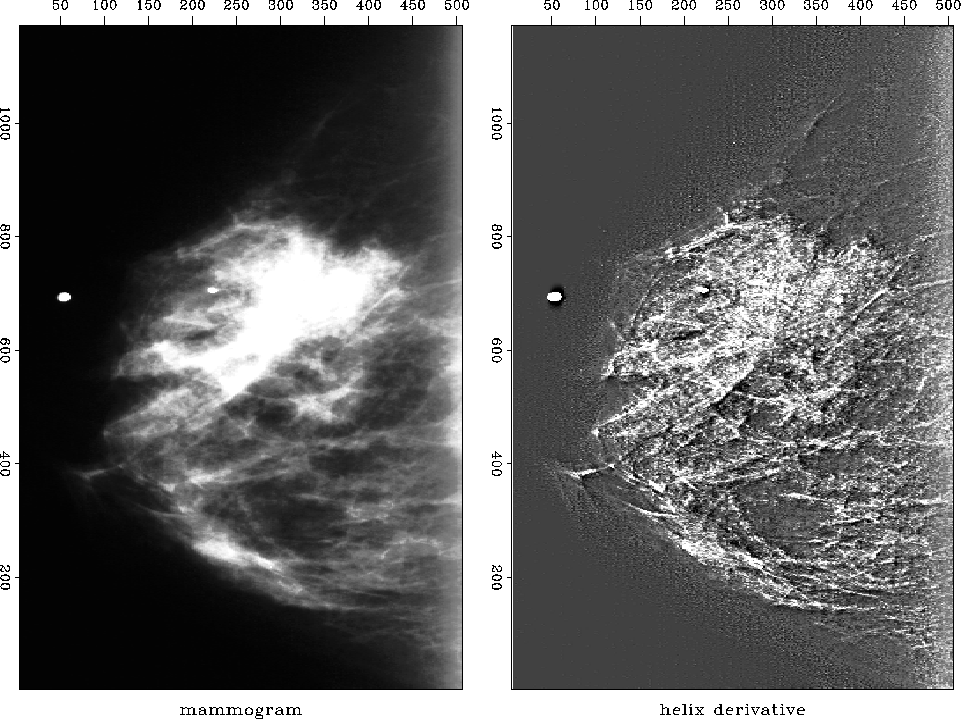 |
![[*]](http://sepwww.stanford.edu/latex2html/movie.gif)