|
rabdomain
Figure 10 Domain of inputs and outputs of a two-dimensional filter like a PEF. | 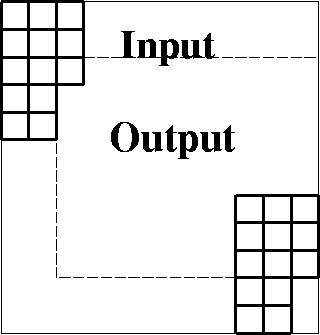 |
Sometimes we deal with small patches of data. In order that boundary phenomena not dominate the calculation intended in the central region, we need to take care that input data is not assumed to be zero beyond the interval that the data is given.
The two little triangular patches of zeros in the convolution matrix in equation (27) describe end conditions where it is assumed that the data yt vanishes before t=1 and after t=6. Alternately we might not wish to make that assumption. Thus the triangles filled with zeros could be regarded as missing data. In this one-dimensional example, it is easy to see that the filter, say yy%mis() should be set to .TRUE. at the ends so no output would ever be computed there. We would like to find a general multidimensional algorithm to correctly specify yy%mis() around the multidimensional boundaries. This proceeds like the missing data algorithm, i.e. we apply a filter of all ones to a data space template that is taken all zeros except ones at the locations of missing data, in this case y0,y-1 and y7,y8. This amounts to surrounding the original data set with some missing data. We need padding the size of the filter on all sides. The padded region would be filled with ones (designating missing inputs). Where the convolution output is nonzero, there yy%mis() is set to .TRUE. denoting an output with missing inputs.
The two-dimensional case is a little more cluttered than the 1-D case but the principle is about the same. Figure 10 shows a larger input domain, a filter, and a smaller output domain.
|
rabdomain
Figure 10 Domain of inputs and outputs of a two-dimensional filter like a PEF. |  |
There are two things to notice. First, sliding the filter everywhere inside the outer box, we get outputs (under the 1 location) only in the inner box. Second, (the adjoint idea) crosscorrelating the inner and outer boxes gives us the patch of information we use to build the filter coefficients. We need to be careful not to assume that signals vanish outside the region where they are defined. In a later chapter we will break data spaces into overlapping patches, separately analyze the patches, and put everything back together. We do this because crosscorrelations change with time and they are handled as constant in short time windows. There we must be particularly careful that zero signal values not be presumed outside of the small volumes; otherwise the many edges and faces of the many small volumes can overwhelm the interior that we want to study.
In practice, the input and output are allocated equal memory,
but the output residual is initialized to zero everywhere
and then not computed
except where shown in figure 10.
Below is module bound
to build a selector for filter outputs that should
never be examined or even computed
(because they need input data from outside the given data space).
Inputs are a filter aa
and the size of its cube na = (na(1),na(2),...).
Also input are two cube dimensions,
that of the data last used by the filter nold and
that of the filter's next intended use nd.
(nold and nd are often the same).
Module bound
begins by defining a bigger data space with room for a filter
surrounding the original data space nd on all sides.
It does this by the line nb=nd+2*na.
Then we allocate two data spaces
xx and yy of the bigger size nb
and pack many ones
in a frame of width na around the outside of xx.
The filter aa is also filled with ones.
The filter aa must be regridded for the bigger nb
data space (regridding merely changes the lag values of the ones).
Now we filter the input xx with aa getting yy.
Wherever the output is nonzero,
we have an output that has been affected by the boundary.
Such an output should not be computed.
Thus we allocate the logical mask aa%mis
(a part of the helix filter definition
in module helix
module bound { # mark helix filter outputs where input is off data.
use cartesian
use helicon
use regrid
contains
subroutine boundn ( nold, nd, na, aa) {
integer, dimension( :), intent( in) :: nold, nd, na # (ndim)
type( filter) :: aa
integer, dimension( size( nd)) :: nb, ii
real, dimension( :), allocatable :: xx, yy
integer :: iy, my, ib, mb, stat
nb = nd + 2*na; mb = product( nb) # nb is a bigger space to pad into.
allocate( xx( mb), yy( mb)) # two large spaces, equal size
xx = 0. # zeros
do ib = 1, mb { # surround the zeros with many ones
call line2cart( nb, ib, ii) # ii( ib)
if( any( ii <= na .or. ii > nb-na)) xx( ib) = 1.
}
call helicon_init( aa) # give aa pointer to helicon.lop
call regridn( nold, nb, aa); aa%flt = 1. # put all 1's in filter
stat = helicon_lop( .false., .false., xx, yy) # apply filter
call regridn( nb, nd, aa); aa%flt = 0. # remake filter for orig data.
my = product( nd)
allocate( aa%mis( my)) # attach missing designation to y_filter
do iy = 1, my { # map from unpadded to padded space
call line2cart( nd, iy, ii )
call cart2line( nb, ii+na, ib ) # ib( iy)
aa%mis( iy) = ( yy( ib) > 0.) # true where inputs missing
}
deallocate( xx, yy)
}
}
![[*]](http://sepwww.stanford.edu/latex2html/cross_ref_motif.gif) )
and wherever we see a nonzero value of yy
in the output,
we designate the output as depending on missing inputs by setting
aa%mis to .true..
)
and wherever we see a nonzero value of yy
in the output,
we designate the output as depending on missing inputs by setting
aa%mis to .true..
#$
#$=head1 NAME
#$
#$bound - find the boundaries of a filter on given map
#$
#$=head1 SYNOPSIS
#$
#$C<call boundn(nold,nd,na,aa)>
#$
#$=head1 INPUT PARAMETERS
#$
#$=over 4
#$
#$=item nold - C<int(:)>
#$
#$ ?????
#$
#$=item nd - C<int(:)>
#$
#$ Size of data
#$
#$=item na - int(:)
#$
#$ Size of box surrounding filter
#$
#$=item aa - C<type(filter)>
#$
#$ Filter to calculate boundaries for
#$
#$
#$=back
#$
#$=head1 DESCRIPTION
#$
#$mark helix filter outputs where input is off data
#$
#$=head1 SEE ALSO
#$
#$L<cartesian>,L<nbound>,L<helicon>,L<regrid>,L<helix>
#$
#$=head1 LIBRARY
#$
#$B<geef90>
#$
#$=cut
#$
#$
#$
In reality one would set up the boundary conditions with module bound before identifying locations of missing data with module misinput. Both modules are based on the same concept, but the boundaries are more cluttered and confusing which is why we examined them later.